Publications (2000-)
2015- / 2010-2014 / 2005-2009 / 2000-2004
2023
Kudo, F.; Chikuma, T.; Nambu, M.; Chisuga, T.; Sumimoto,S.; Iwasaki, A.; Suenaga,K.; Miyanaga,A.; EguchiT. Unique Initiation and Termination Mechanisms Involved in the Biosynthesis of a Hybrid Polyketide-nonribosomal Peptide Lyngbyapeptin B Produced by the Marine Cyanobacterium Moorena bouillonii. ACS Chem. Biol., in press.
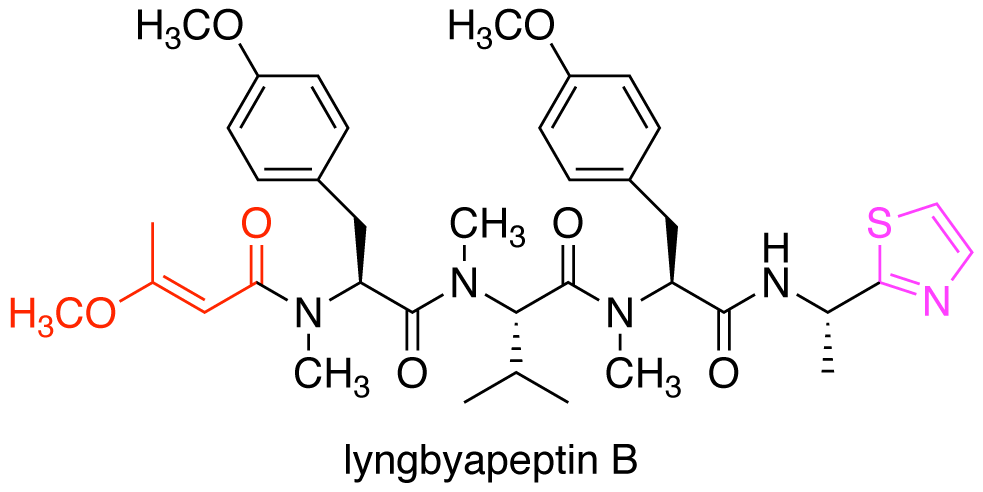
Kudo, F.; Kishikawa, K.; Tsuboi, K.; Kido, T.; Usui,T.; Hashimoto,J.; Shin-ya,K.; Miyanaga, A.; Eguch, T. Acyltransferase Domain Exchanges between Two Independent Type I Polyketide Synthases in the Same Producer Strain of Macrolide Antibiotics. ChemBioChem, 2023, 24, e202200670.
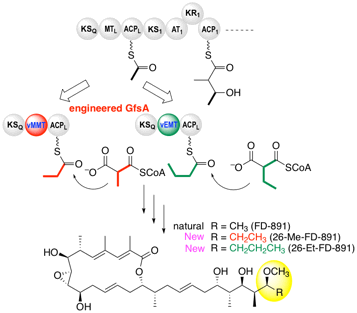
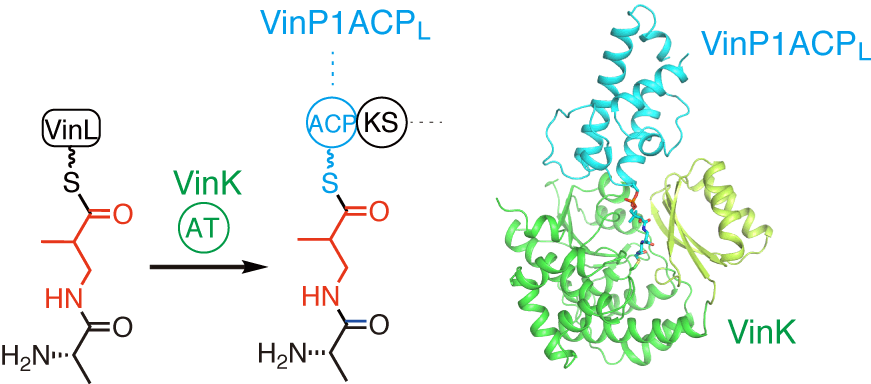
Miyanaga, A.; Kawada, K.; Chisuga, T.; Kudo, F.; Eguchi, T. Structural Basis of Transient Interactions of the Acyltransferase VinK with the Loading Acyl Carrier Protein of the Vicenistatin Modular Polyketide Synthase. Biochemistry, 2023, 62, 17-21.
Watanabe, I.; Miyanaga, A.; Hoshi, H.; Suzukui, K.; Eguchi, T. Biochemical and Crystallographic Assessments of the Effect of 4,6-O-Disulfated Disaccharide Moieties in Chondroitin Sulfate E on Chondroitinase ABC I Activity. FEBS J., in press.
2022
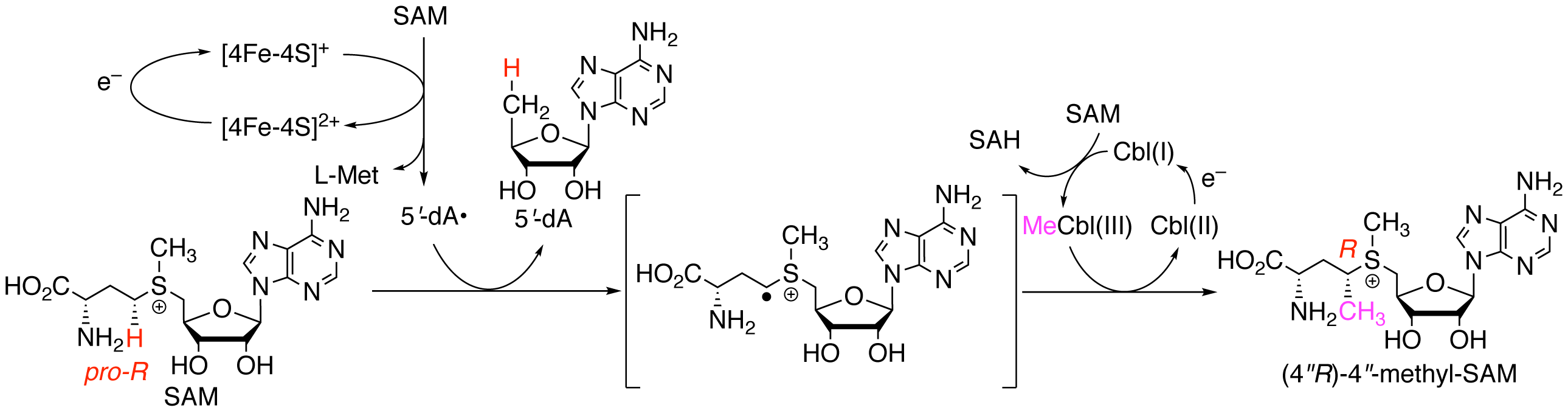
Kudo, F.; Minato, A.; Sato, S.; Nagano, N.; Maruyama, C.; Hamano, Y.; Hashimoto, J.; Kozone, I.; Shin-ya, K.; Eguchi, T. Mechanism of S-Adenosyl-L-methionine C-Methylation by Cobalamin-dependent Radical S-Adenosyl-L-methionine Methylase in 1-Amino-2-methylcyclopropanecarboxylic Acid Biosynthesis. Org. Lett., 2022, 24, 8975-8979.
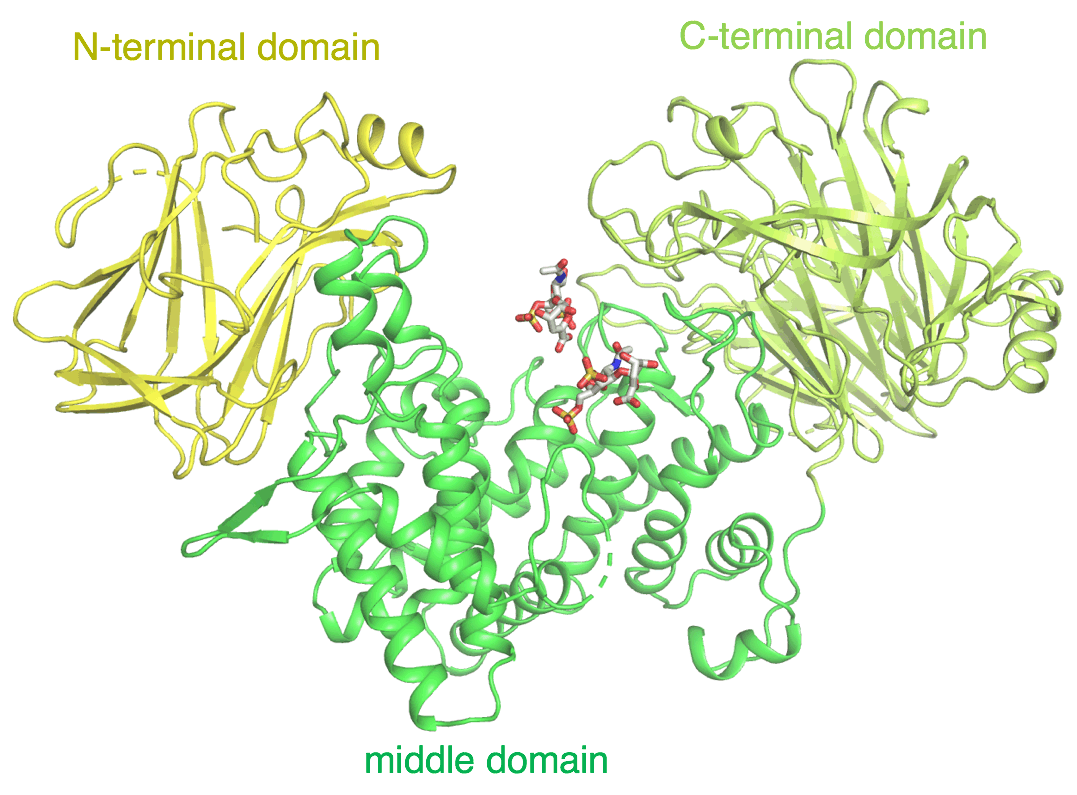
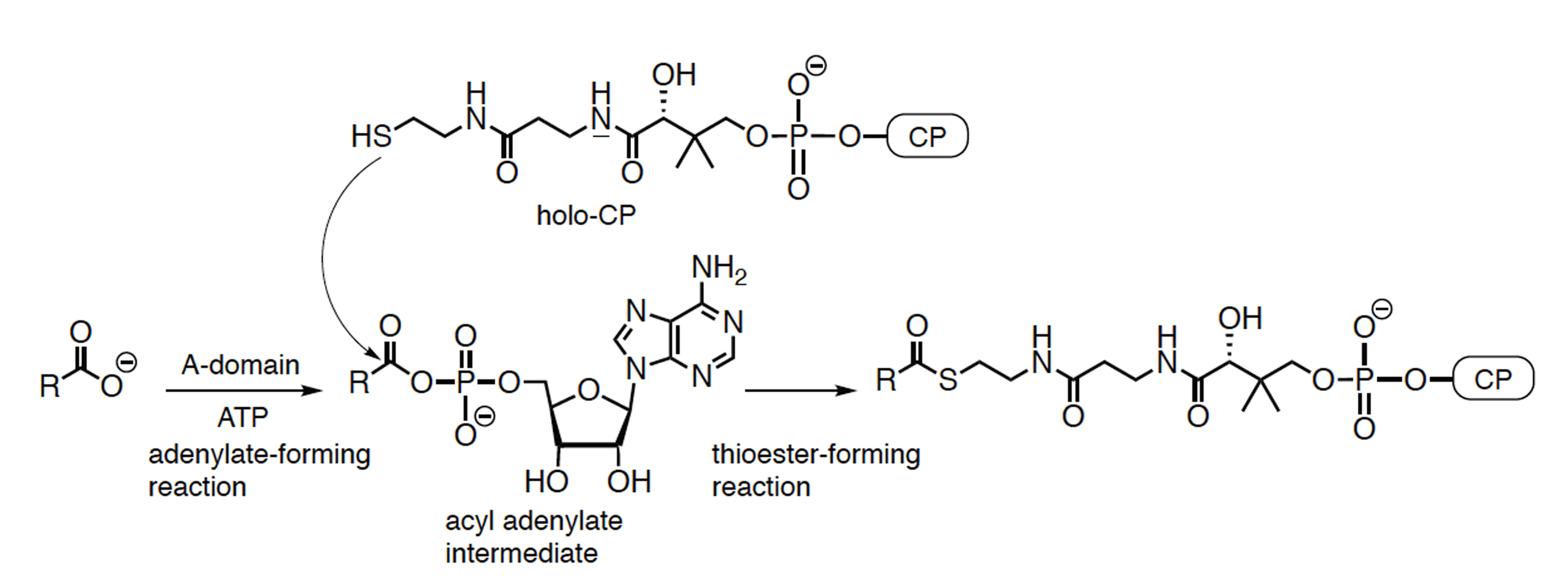
Miyanaga, A.; Kudo, F.; Eguchi, T. Recent Advances in Structural Analysis of Adenylation Domains in Natural Product Biosynthesis. Curr. Opin. Chem. Biol., 2022, 71, e102212.
. 
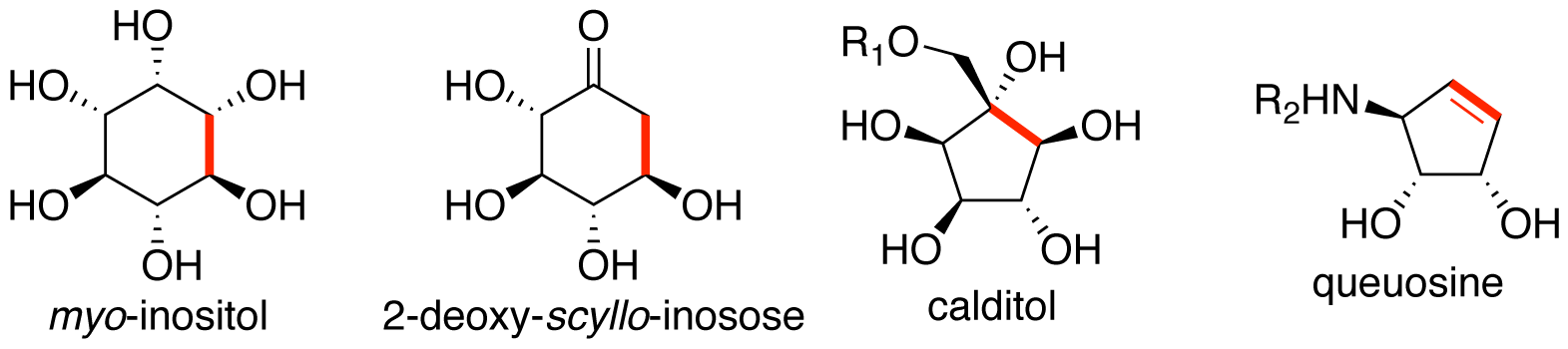
Kudo, F.; Eguchi, T. Biosynthesis of Cyclitols. Nat. Prod. Rep., 2022, 39, 1622-1642.
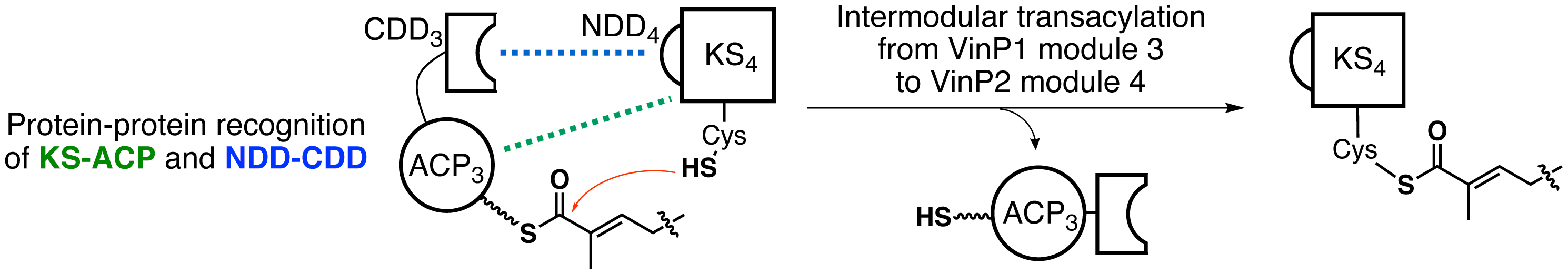
Chisuga, T.; Miyanaga, A.; Eguchi, T. Protein–protein Recognition Involved in Intermodular Transacylation Reaction in Modular Polyketide Synthase in the Biosynthesis of Vicenistatin. ChemBioChem, 2022, 23, e202200200.
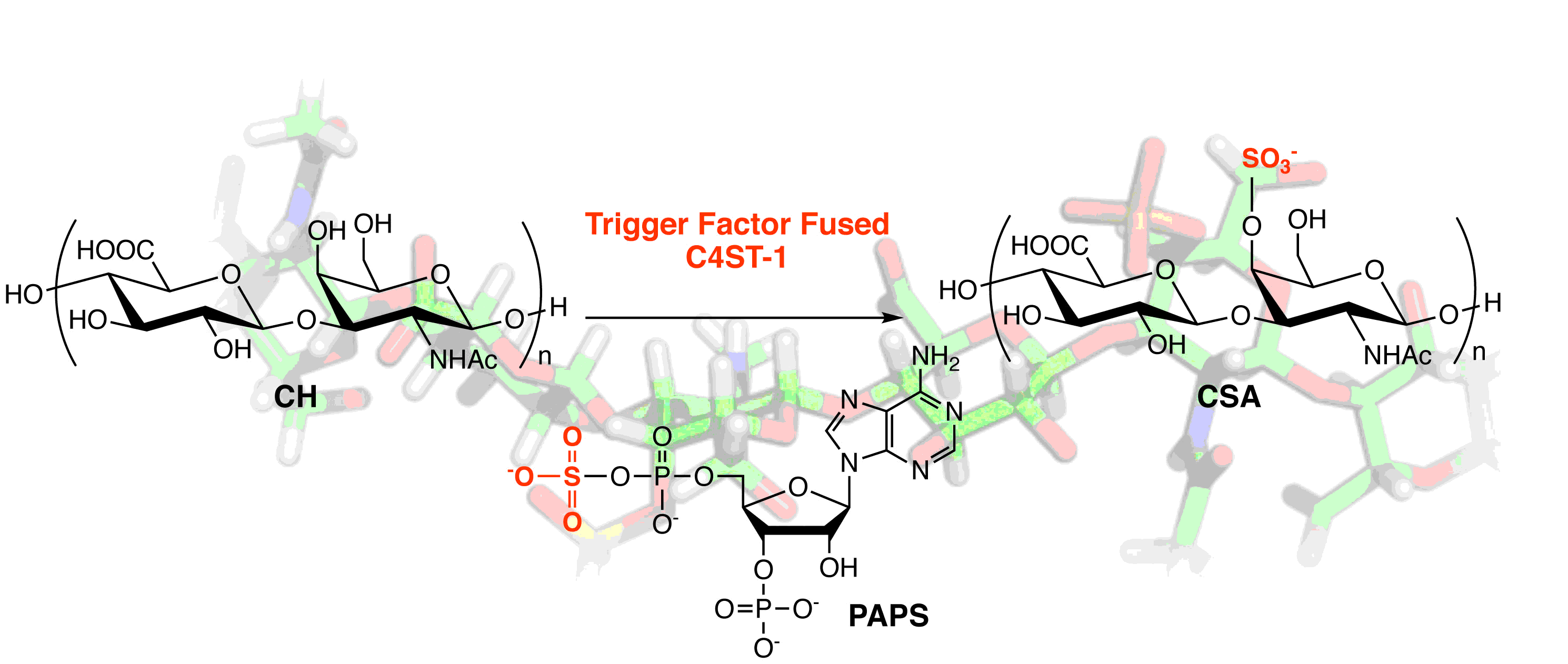
Takashima, M.; Suzuki, K.; Mochizuki, H.; Uemura, S.; Inokuchi, J.; Eguchi, T. Expression of Highly Active Chondroitin 4-O-Sulfotransferase-1 in Escherichia coli by a Trigger Factor Fusion Protein Expression System. Process Biochem., 2022, 115, 146-151.
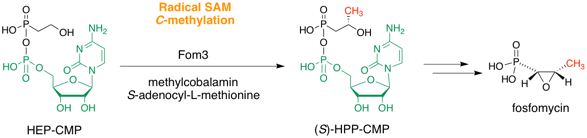
Sato, S.; Kudo, F.; Eguchi, T. Charcterization of the Cobalamin-Dependent Radical SAM C-Methyltransferase Fom3 in Fosfomycin Biosynthesis. Methods Enzymol., 2022, 669, 45-70.
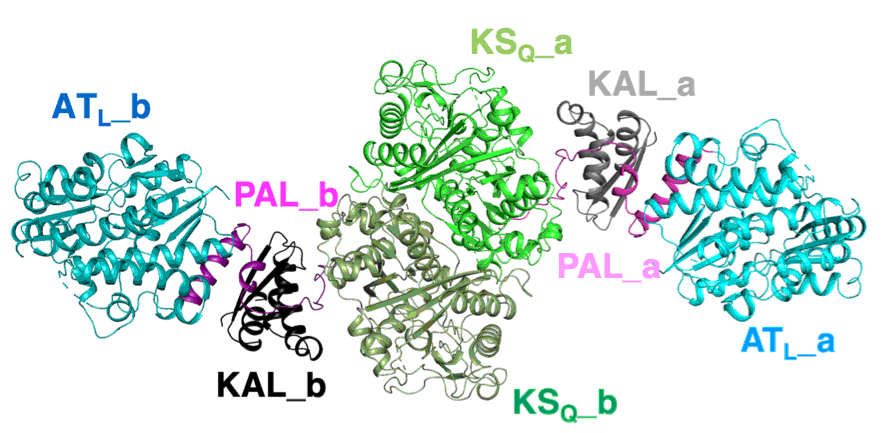
Chisuga, T.; Nagai, A.; Miyanaga,A.; Goto, E.; Kishikawa, K.; Kudo, F.; Eguchi, T. Structural Insight into the Reaction Mechanism of Ketosynthase-like Decarboxylase in a Loading Module of Modular Polyketide Synthases. ACS Chem. Biol., 2022, 17, 198-206.
2021
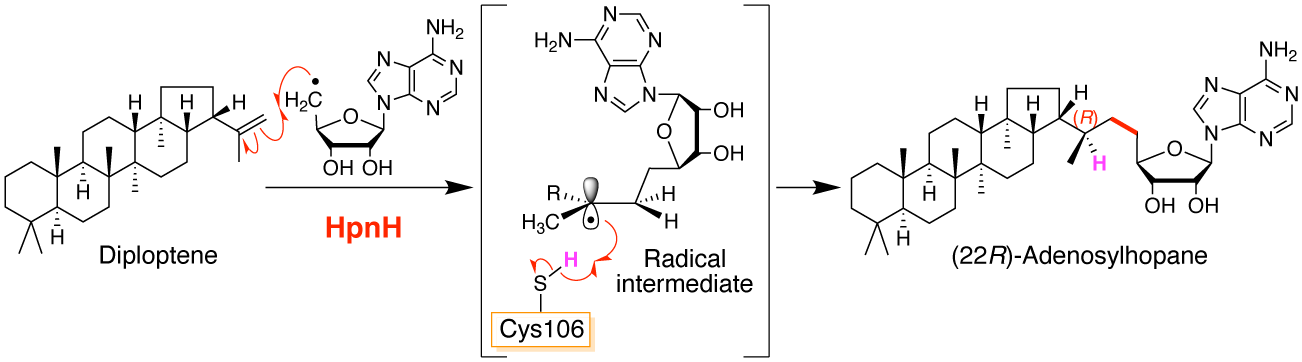
Sato, S.; Kudo, F.; Rohmer, M.; Eguchi, T. Biochemical and Mutational Analysis of Radical SAM Adenosylhopane Synthase HpnH from Zymomonas mobilis Reveals that the Conserved Residue Cysteine-106 Reduces a Radical Intermediate and Determines the Stereochemistry. Biochemistry, 2021, 60, 2865-2874.
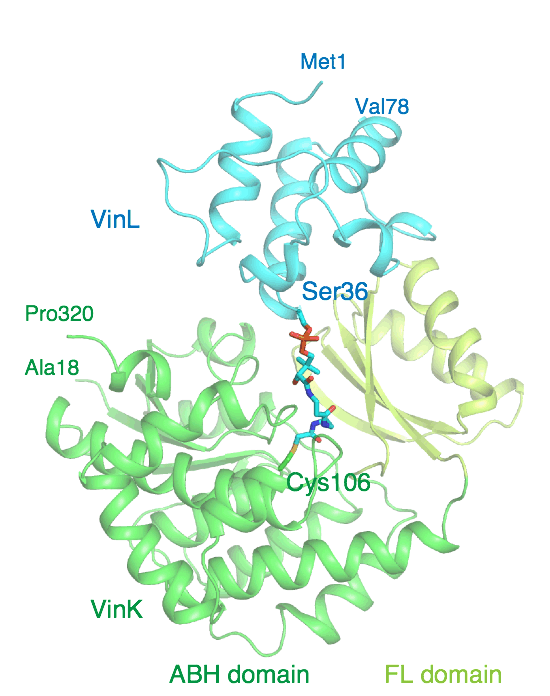
Miyanaga, A.; Ouchi, R.; Kudo, F.; Eguchi, T. Complex Structure of the Acyltransferase VinK and the Carrier Protein VinL with a Pantetheine Cross-linking Probe. Acta Crystallogr F Struct. Biol. Commun., 2021, F77, 294-302.
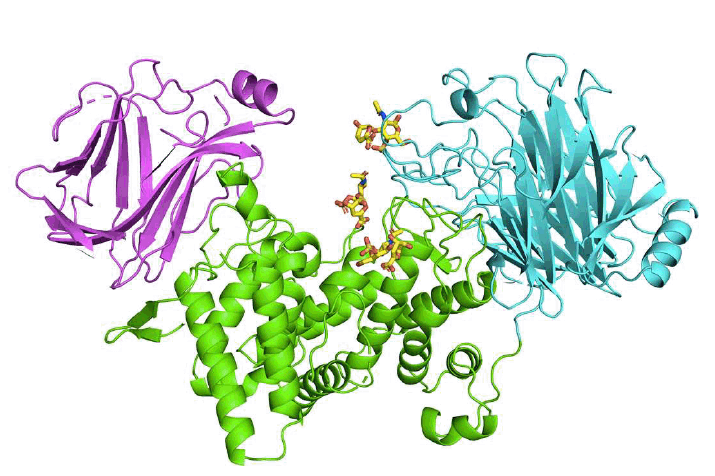
Takashima, M.; Watanabe, I.; Miyanaga, A.; Eguchi, T. Substrate Specificity of Chondroitinase ABC I Based on Analyses of Biochemical Reactions and Crystal Structures in Complex with Disaccharides. Glycobiology, 2021, 31, 1571-1581.
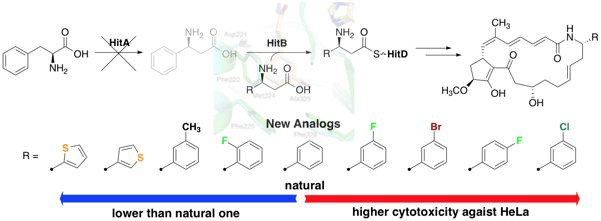
Kudo, F.; Takahashi, S.; Miyanaga, A.; Nakazawa, Y.; Nishino, K.; Hayakawa, Y.; Kawamura, K.; Ishikawa, F.; Tanabe, G.; Iwai, N.; Nagumo, Y.; Usui, T.; Eguchi, T. Mutational Biosynthesis of Hitachimycin Analogs Controlled by the β-Amino Acid–Selective Adenylation Enzyme HitB. ACS Chem. Biol., 2021, 16, 539-547.
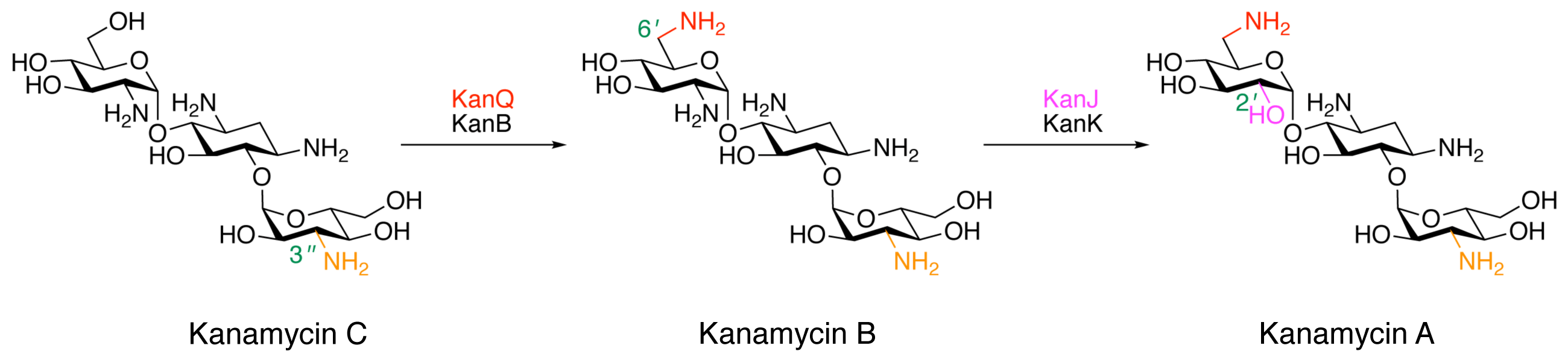
Kudo, F.; Kitayama, Y.;: Miyanaga, A.; Numakura, M.; Eguchi, T. Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis. ChemBioChem, 2021, 22, 1668-1675.
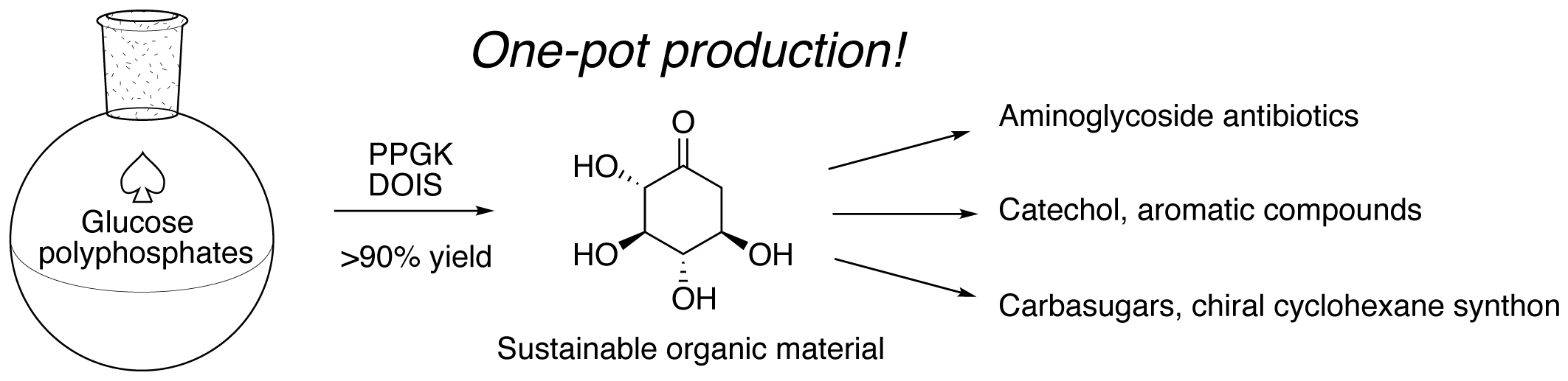
Kudo, F.; Mori, A.; Koide, M.; Yajima, R.; Takeishi,T.; Miyanaga, A.; Eguchi, T. One-pot Enzymatic Synthesis of 2-Deoxy-scyllo-inosose from D-Glucose and Polyphosphate. Biosci. Biotechnol. Biochem., 2021, 85, 108-114.
2020
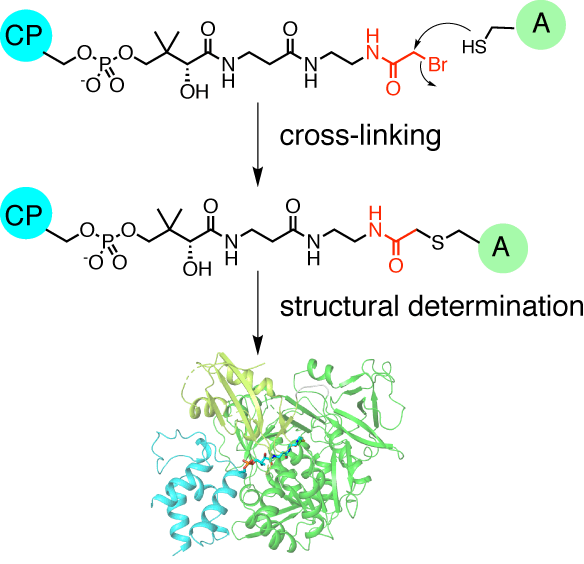
Miyanaga, A.; Kurihara, S.; Chisuga, T.; Kudo, F.; Eguchi, T. Structural Characterization of Complex of Adenylation Domain and Carrier Protein by using Pantetheine Cross-linking Probe. ACS Chem. Biol., 2020, 15, 1808-1812.
Miyanaga, A.; Takaku,R.; Takaishi, M.; Tashiro, E.; Kudo, F.; Eguchi, T. Generation of Incednine Derivatives by Mutasynthesis. J. Antibiot., 2020, 73, 794-797.

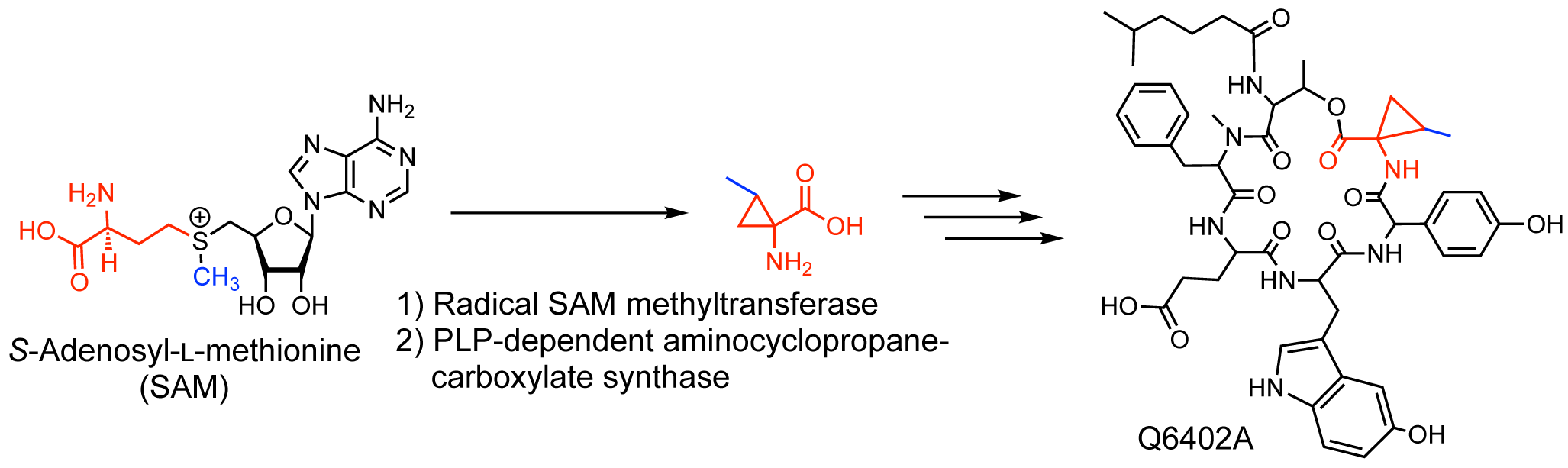
Maruyama, C.; Chinone, Y.; Sato, S.; Kudo, F.; Ohsawa, K.; Kubota, J.; Hashimoto, J.; Kozone, I.; Doi, T.; Shin-ya, K.; Eguchi, T.; Hamano Y.
C-Methylation of S-Adenosyl-L-methionine Occurs prior to Cyclopropanation in the Biosynthesis of a Bacterial 1-Amino-2-methylcyclopropanecarboxylic Acid (Norcoronamic acid) in a Bacterium. Biomolecules, 2020, 10, E775 (2020).
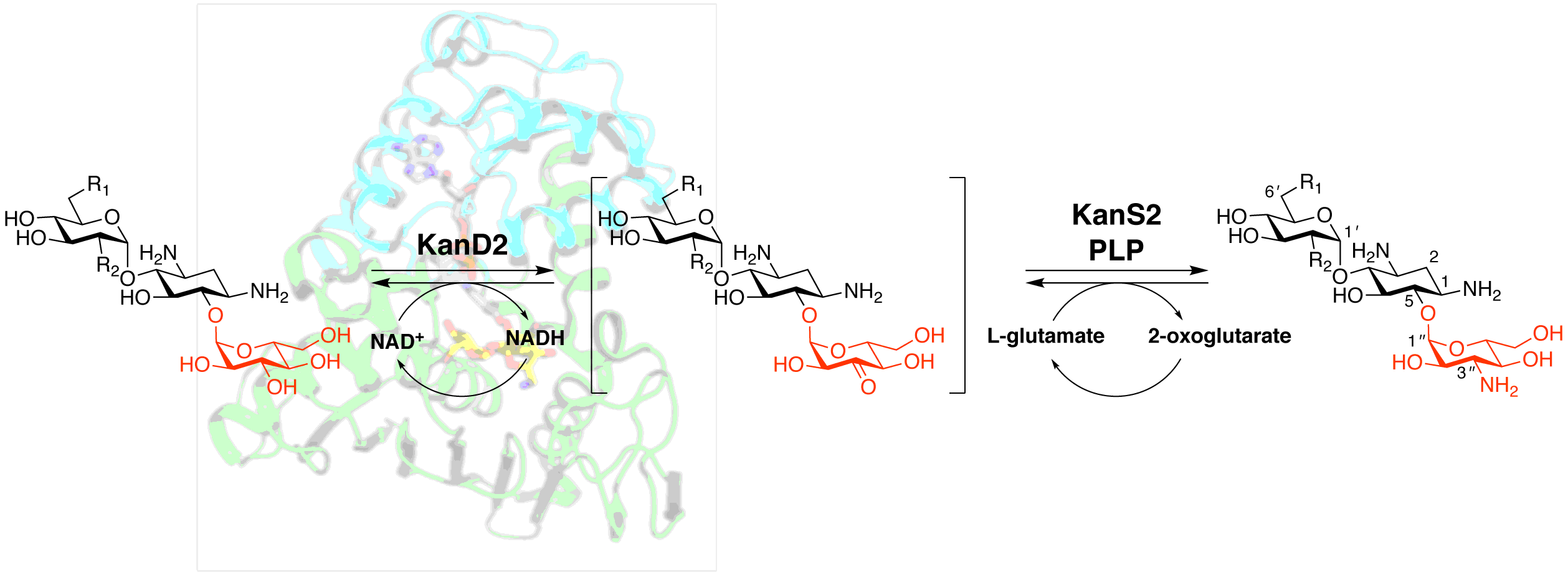
Kudo, F.; Kitayama, Y.; Miyanaga, A.; Hirayama, A.; Eguchi T. Biochemical and Structural Analysis of a Dehydrogenase KanD2 and an Aminotransferase KanS2 that are Responsible for the Construction of the Kanosamine Moiety in Kanamycin Biosynthesis. Biochemistry, 2020, 59, 1470-1473.
工藤史貴, 江口正, 生体内におけるラジカル反応:ラジカル酵素による炭素–炭素結合形成反応. 月刊「化学」, 2020, 75, 72-73.
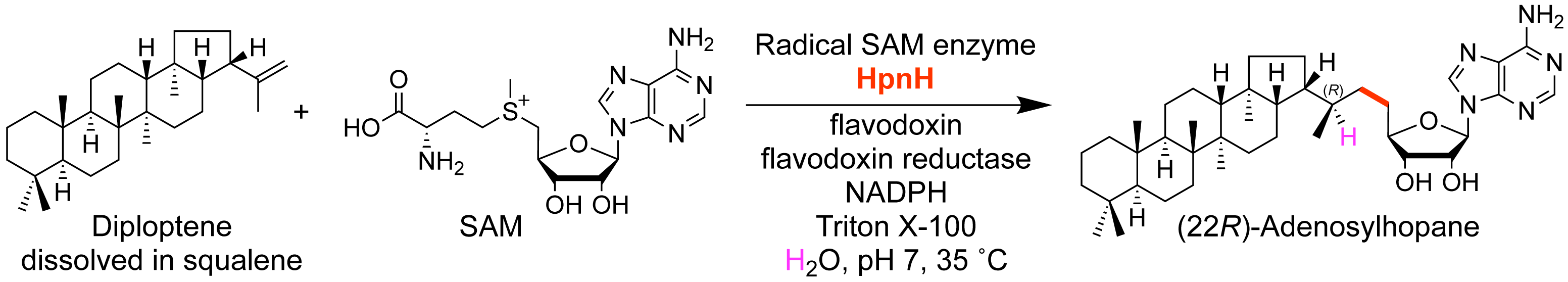
Sato, S.; Kudo, F.; Rohmer, M.; Eguchi, T. Characterization of Radical SAM Adenosylhopane Synthase, HpnH, which Catalyzes the 5'-Deoxyadenosyl Radical Addition to Diploptene in the Biosynthesis of C35 Bacteriohopanepolyols. Angew. Chem. Int. Ed., 2020, 59, 237-241.
2019
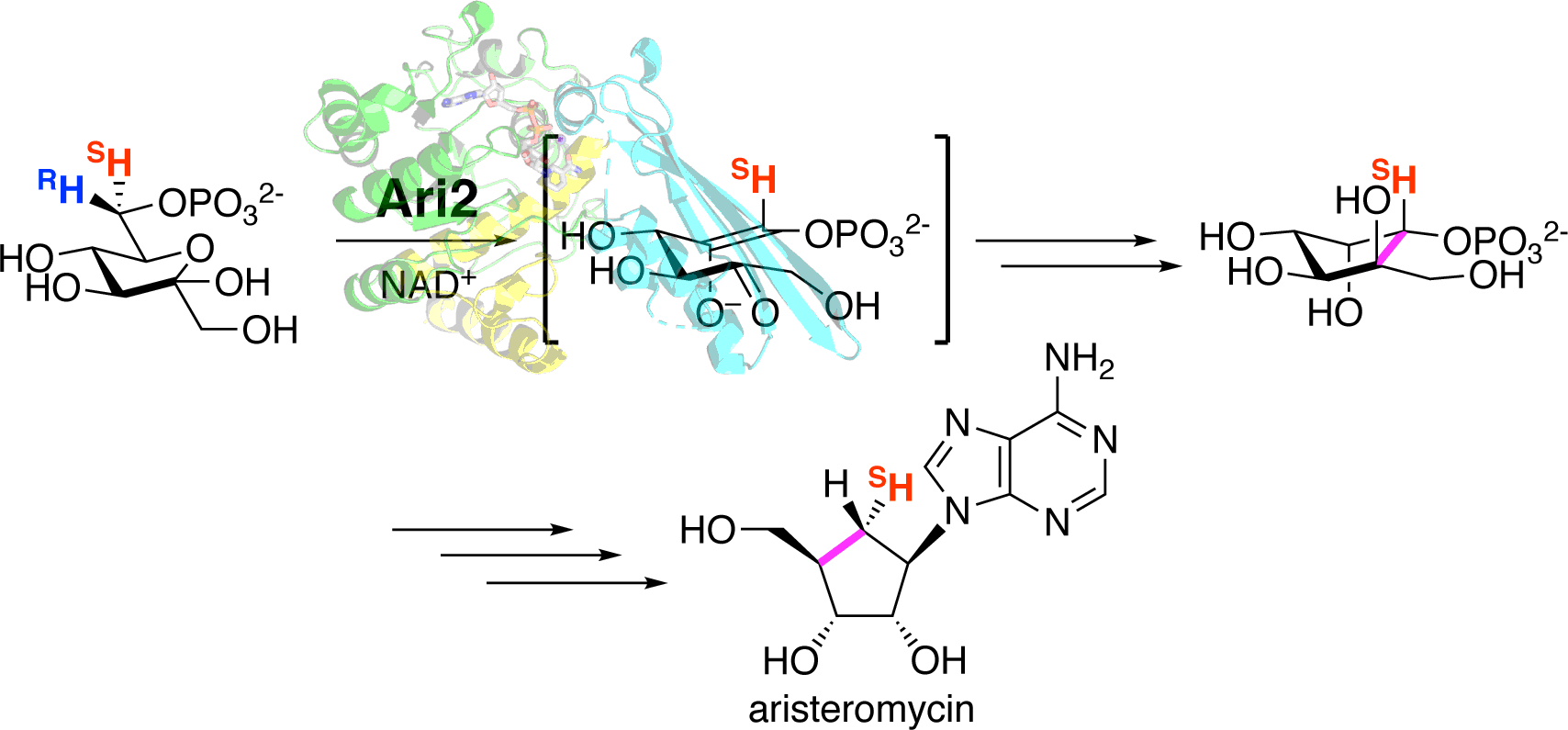
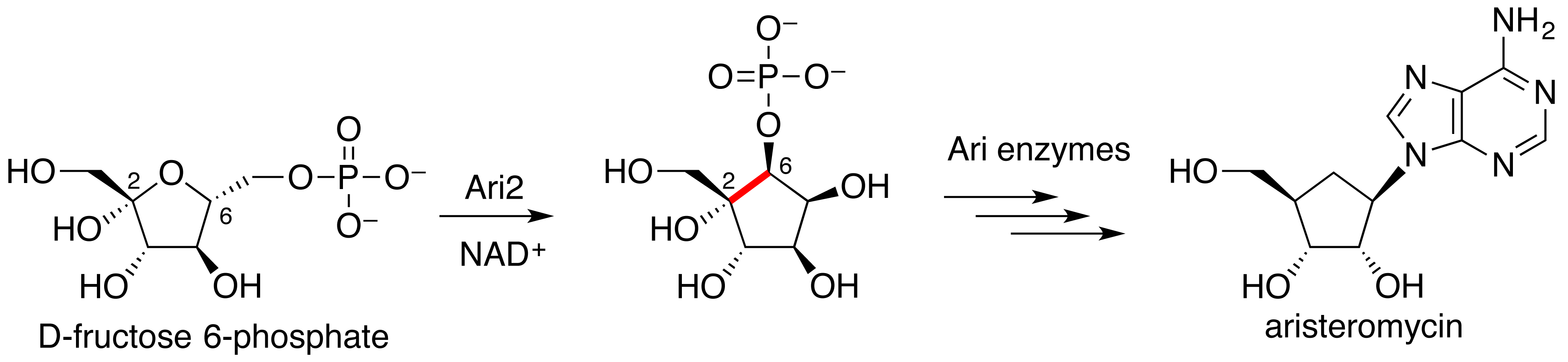
Kudo, F.; Tsunoda T.; Yamaguchi, K.; Miyanaga, A.; Eguchi T. Stereochemistry in the Reaction of the myo-Inositol Phosphate Synthase Ortholog Ari2 during Aristeromycin Biosynthesis. Biochemistry, 2019, 58, 5112-5116.
Kawasaki, D.; Miyanaga, A.; Chisuga, T.; Kudo,F.; Eguchi, T. Functional and Structural Analyses of Split-Dehydratase Domain in the Biosynthesis of Macrolactam Polyketide Cremimycin. Biochemistry, 2019, 58, 4799-4803. 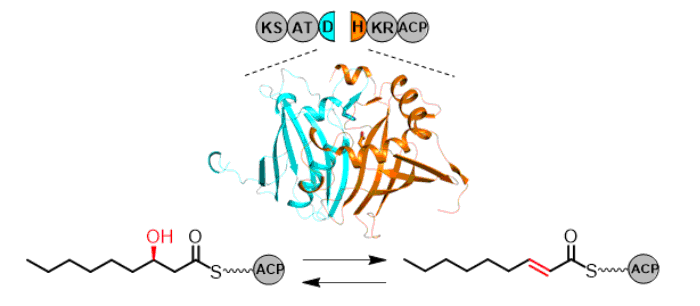
佐藤秀亮, 工藤史貴, 江口正, 抗生物質ホスホマイシン生合成の全貌解明, バイオサイエンスとインダストリー, 2019, 77, 378-379.
Kawasaki, D.; Chisuga, T.; Miyanaga, A.; Kudo, F.; Eguchi, T. Structural Analysis of Glycine Oxidase Homologue CmiS2 Reveals a Unique Substrate Recognition Mechanism for Formation of a β-Amino Acid Starter Unit in Cremimycin Biosynthesis. Biochemistry, 2019, 58, 2706-2709.
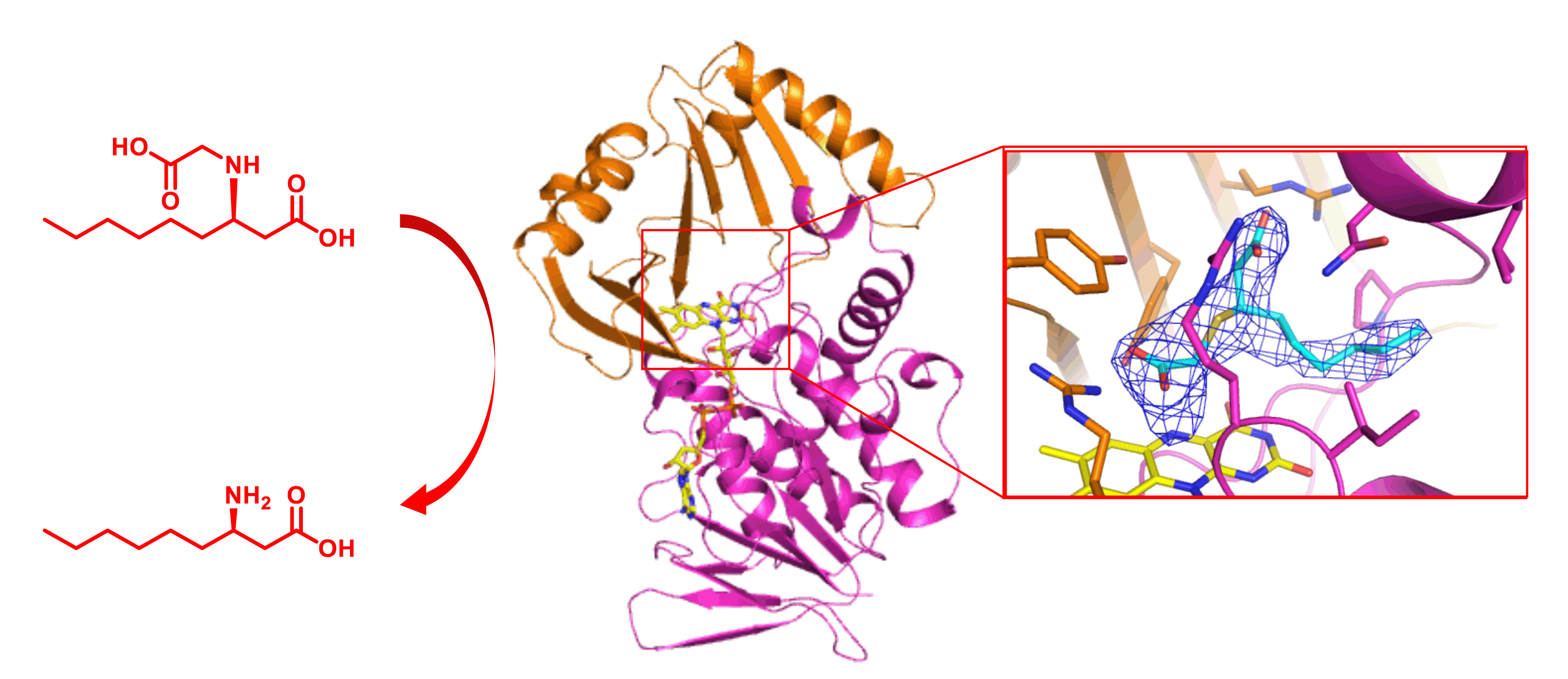
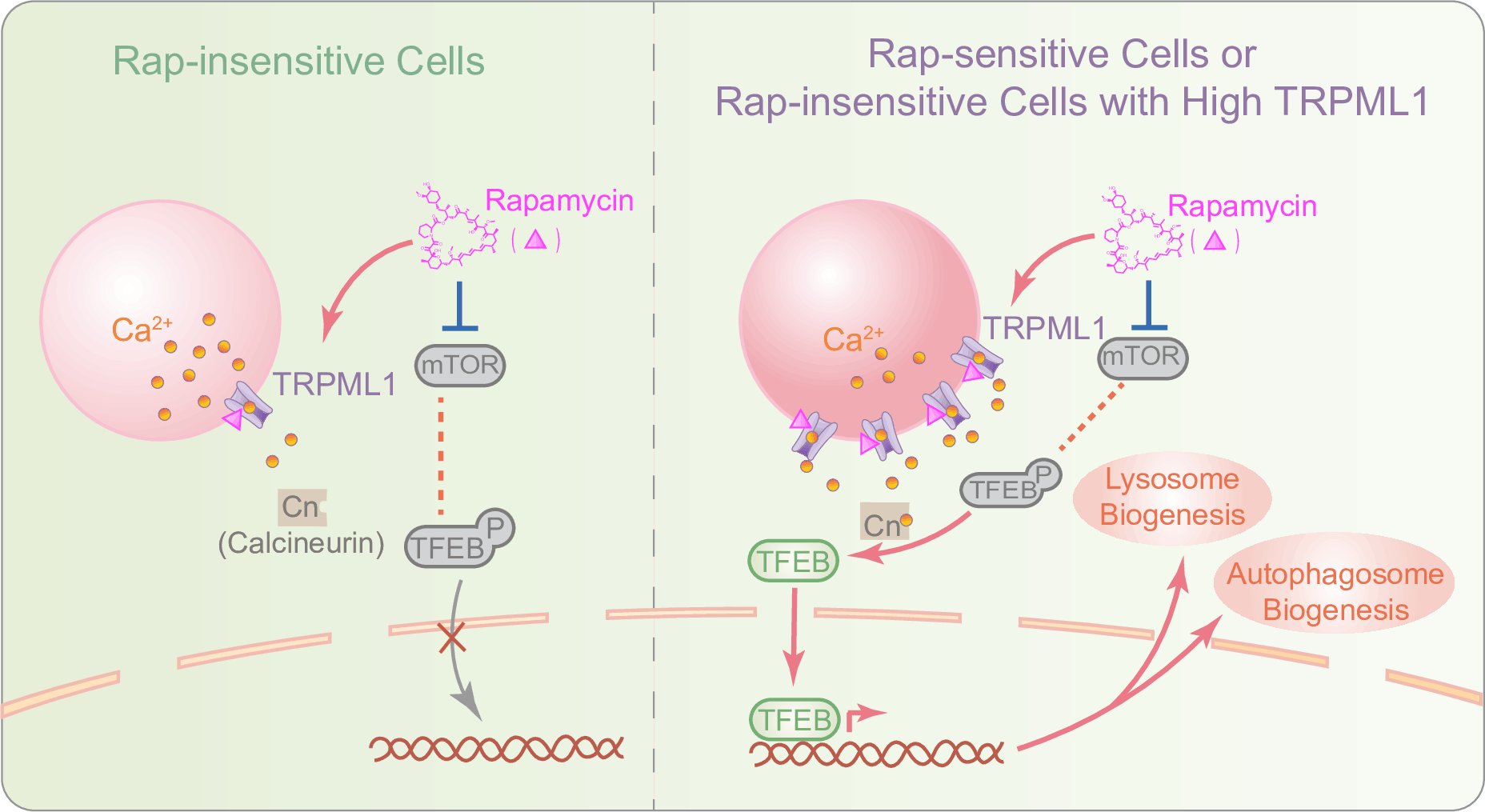
Zhang, X.; Chen, W.; Gao, Q.; Yang, J.; Yan, X.; Zhao, H.; Su, L.; Yang, M.; Gao, C.; Yao, Y.; Inoki, K.; Li, D.; Shao, R.; Wang, S.; Sahoo, N.; Kudo, F.; Eguchi, T.; Ruan,B.; Xu, H. Rapamycin Directly Activates Lysosomal Mucolipin TRP Channels Independent of mTOR. PLOS Biol., 2019, 17, e3000252.
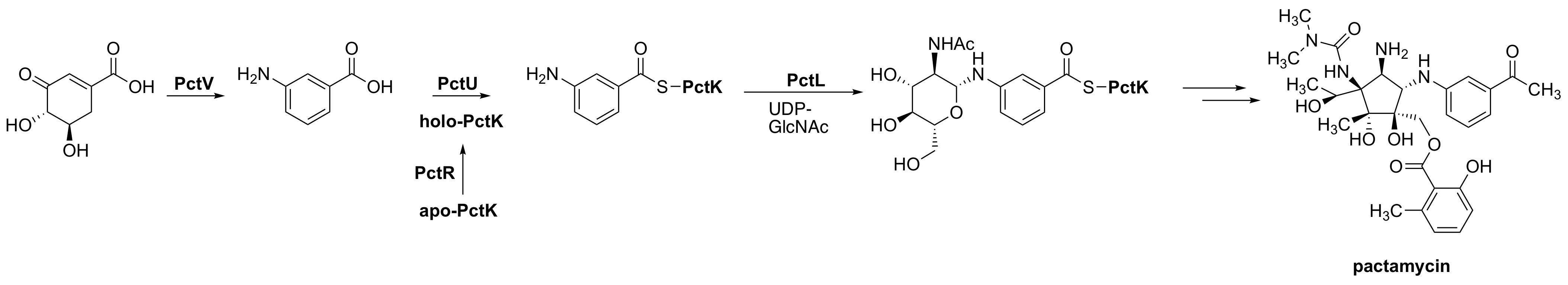
Kudo, F.; Zhang, J.; Sato, S.; Hirayama, A.; Eguchi, T. Functional Characterization of 3-Aminobenzoic Acid Adenylation Enzyme PctU and UDP-N-Acetyl-D-Glucosamine: 3-Aminobenzoyl-ACP Glycosyltransferase PctL in Pactamycin Biosynthesis. ChemBioChem, 2019, 20, 2458-2462.
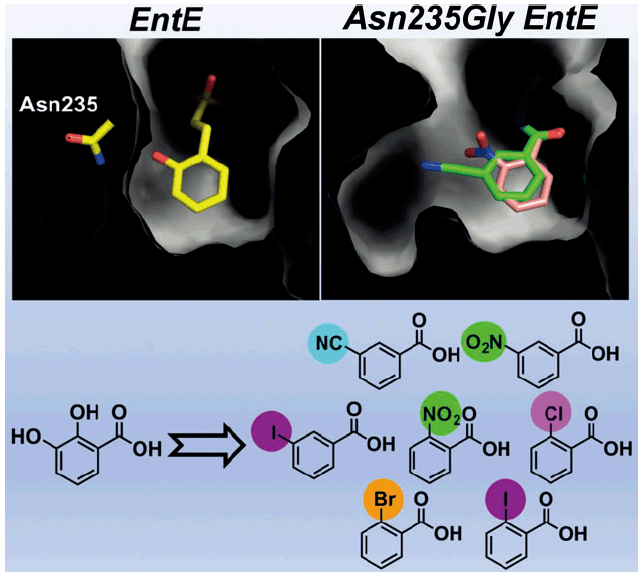
Ishikawa, F.; Miyanaga, A.; Kitayama, H.; Nakamura,S.; Nakanishi,I.; Kudo, F.; Eguchi, T.; Tanabe, G. An Engineered Aryl Acid Adenylation Domain with an Enlarged Substrate Binding Pocket. Angew. Chem. Int. Ed., 2019, 58, 6906-6910.
Cieślak, J.; Miyanaga, A.; Takaishi,T.; Kudo, F.; Eguchi, T. Functional and Structural Characterization of Adenylation Enzyme IdnL7 Involved in Incednine Biosynthesis. Acta Crystallogr. F Struct. Biol. Commun., 2019, F75, 299-306.
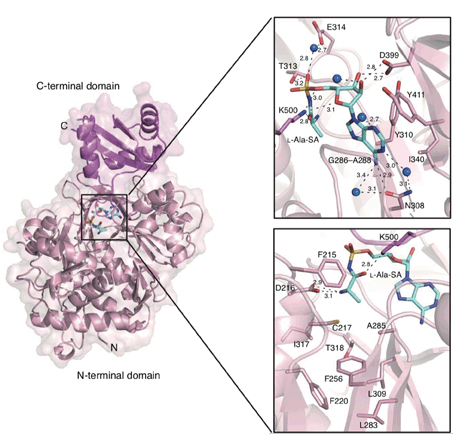
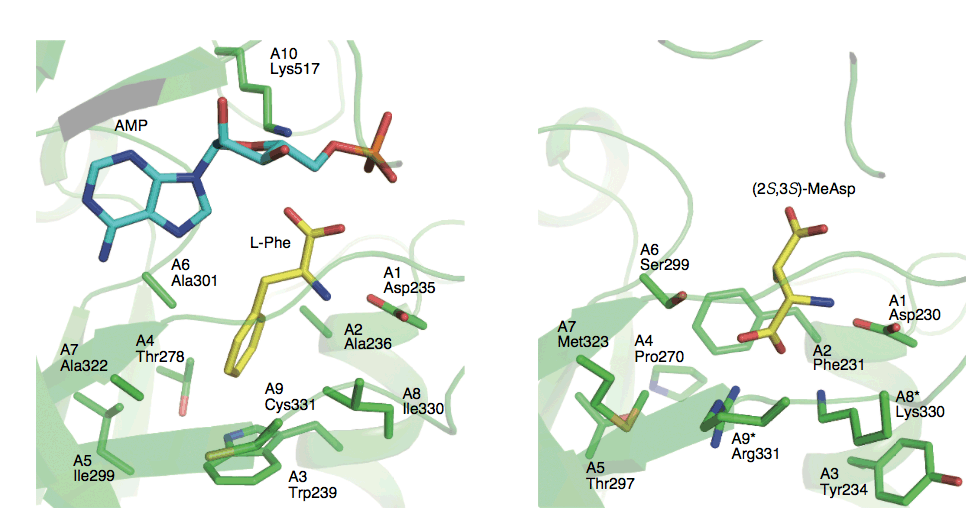
Kudo, F.; Miyanaga, A.; Eguchi, T. Structural Basis of Nonribosomal Codes for Nonproteinogenic Amino Acid Selective Adenylation Enzymes in the Biosynthesis of Natural Products. J. Ind. Microbiol. Biotechnol., 2019, 46, 515-536.
2018
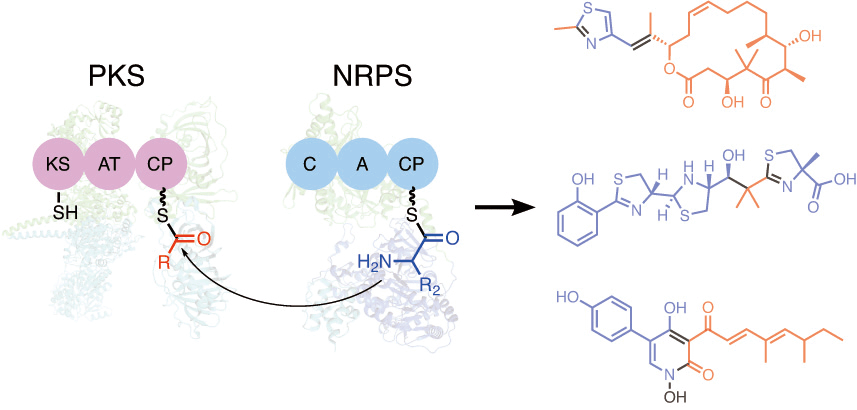
Miyanaga, A.; Kudo, F.; Eguchi, T. Protein–protein Interactions in Polyketide Synthase–Nonribosomal Peptide Synthetase Hybrid Assembly Lines. Nat. Prod. Rep., 2018, 35, 1185-1209.
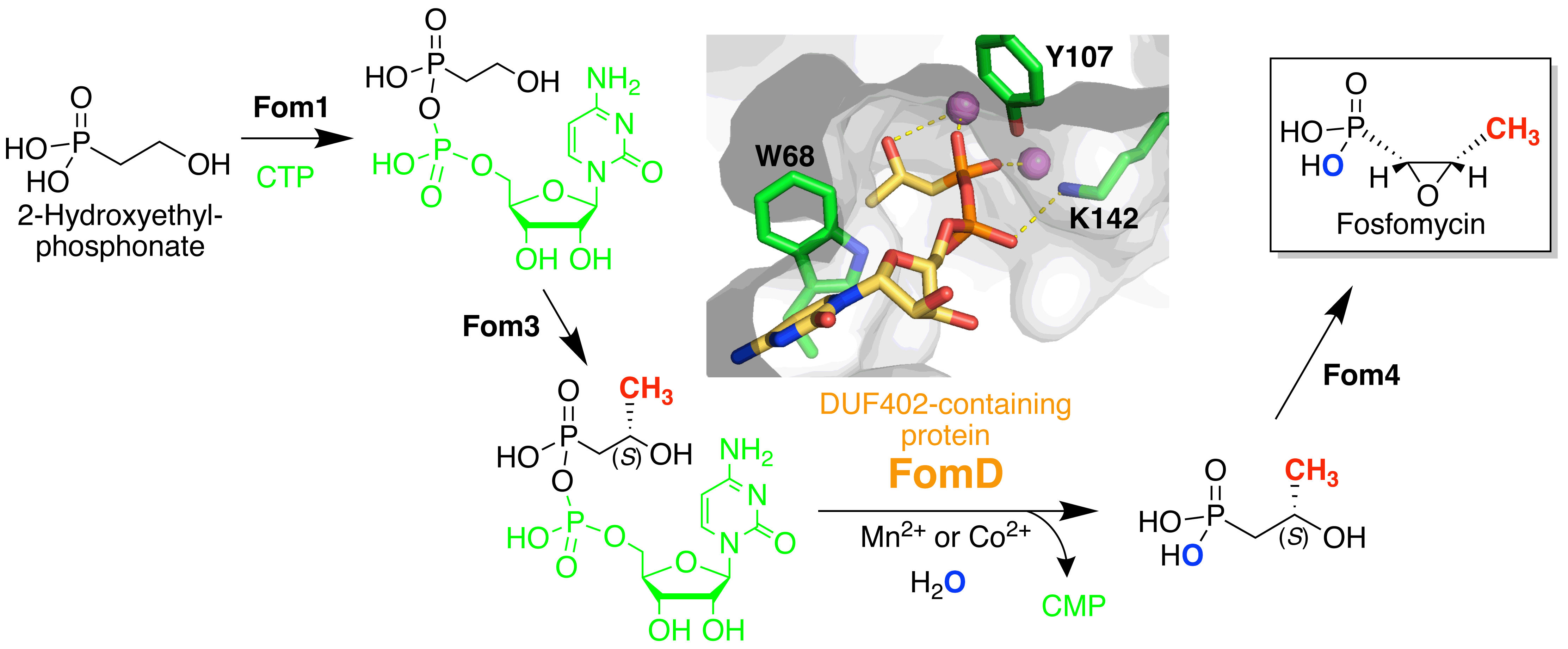
Sato, S.; Miyanaga, A.; Kim, S.-Y.; Kuzuyama, T.; Kudo, F.; Eguchi, T. Biochemical and Structural Analysis of FomD that Catalyzes the Hydrolysis of Cytidylyl (S)-2-Hydroxypropylphosphonate in Fosfomycin Biosynthesis. Biochemistry, 2018, 57, 4858-4866.
Sato, S.; Kudo, K.; Kuzuyama, T.; Hammerschmidt, F.; Eguchi, T. C-Methylation Catalyzed by Fom3, a Cobalamin-Dependent Radical S-Adenosyl-L-methionine Enzyme in Fosfomycin Biosynthesis, Proceeds with Inversion of Configuration. Biochemistry, 2018, 57, 4963-4966.
Miyanaga, A.; Ouchi, R.; Ishikawa, F.; Goto, E.; Tanabe,G.; Kudo, F.; Eguchi, T. Structural Basis of Protein–Protein Interactions between a trans-Acting Acyltransferase and Acyl Carrier Protein in Polyketide Disorazole Biosynthesis. J. Am. Chem. Soc., 2018, 140, 7970-7978.
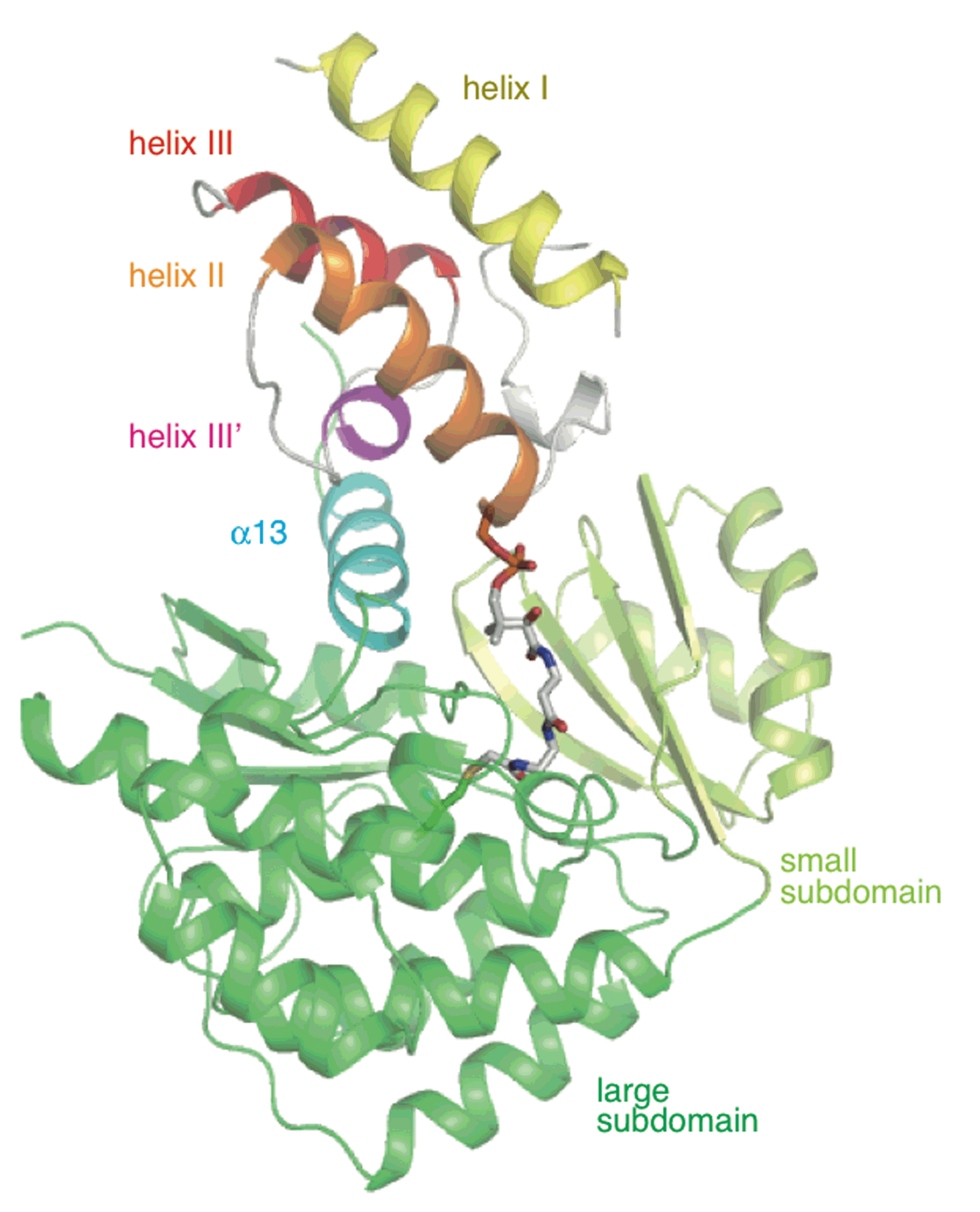
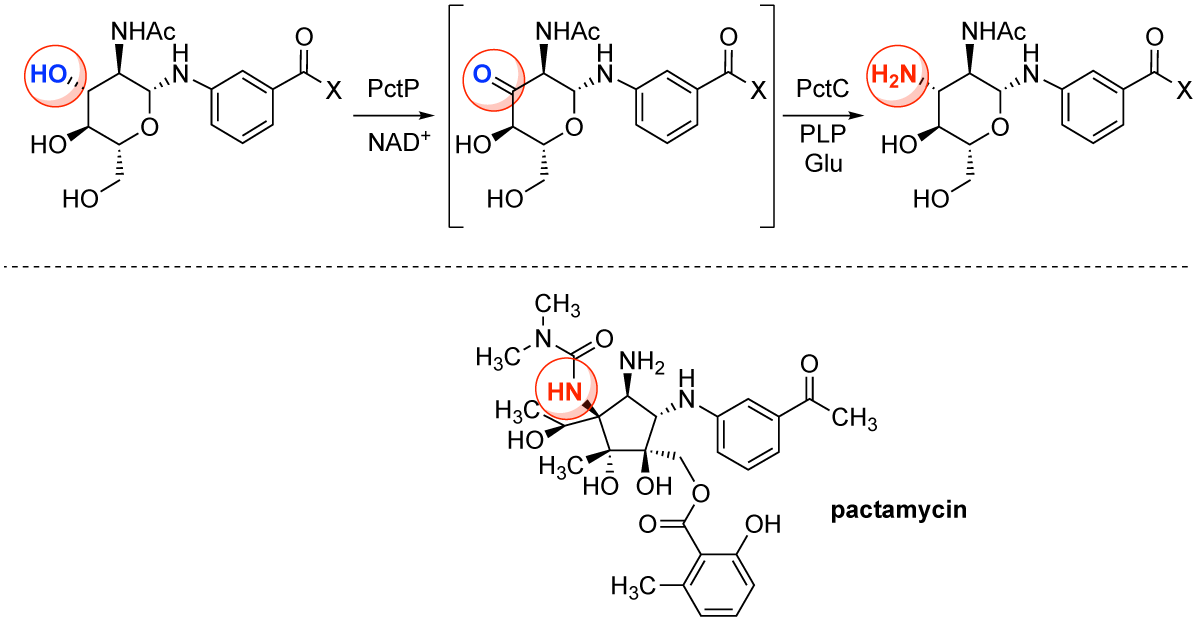
Hirayama, A.; Chu, J.; Goto, E.; Kudo, F.; Eguchi, T. NAD(+)-Dependent Dehydrogenase PctP and PLP-Dependent Aminotransferase PctC Catalyze the First Post-glycosylation Modification of Sugar Intermediate in Pactamycin Biosynthesis. ChemBioChem, 2018, 19, 126-130.
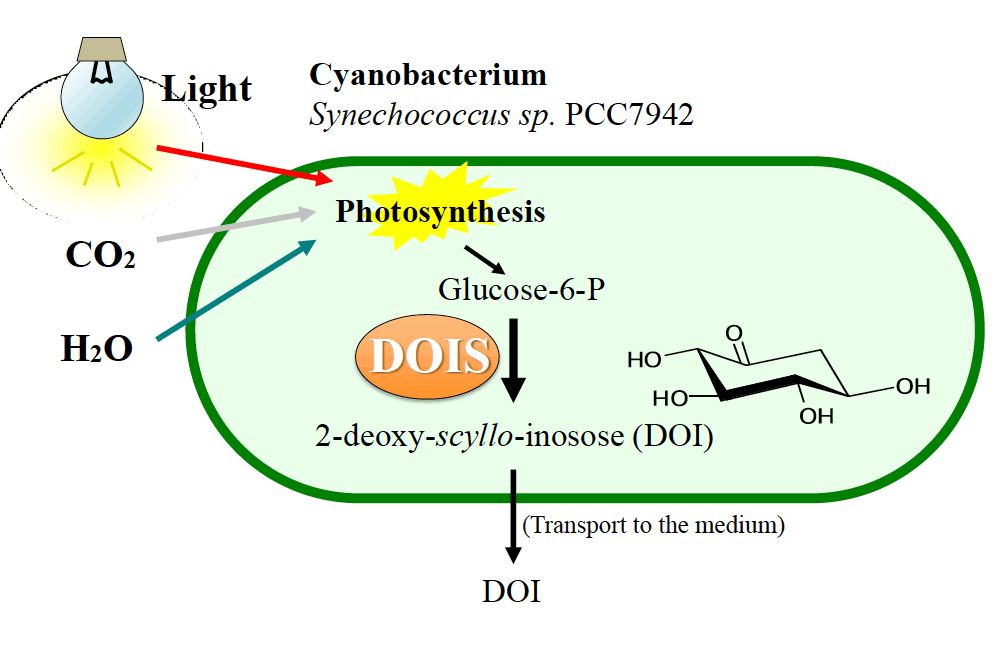
Watanabe, S.; Ozawa, H.; Kato, H.; Nimura-Matsune, K.; Hirayama, T.; Kudo, F.; Eguchi, T.; Kakinuma, K.; Yoshikawa, H. Carbon-free Production of 2-Deoxy-scyllo-inosose (DOI) in Cyanobacterium Synechococcus elongatus PCC 7942. Biosci. Biotechnol. Biochem., 2018, 82, 161-165.
2017
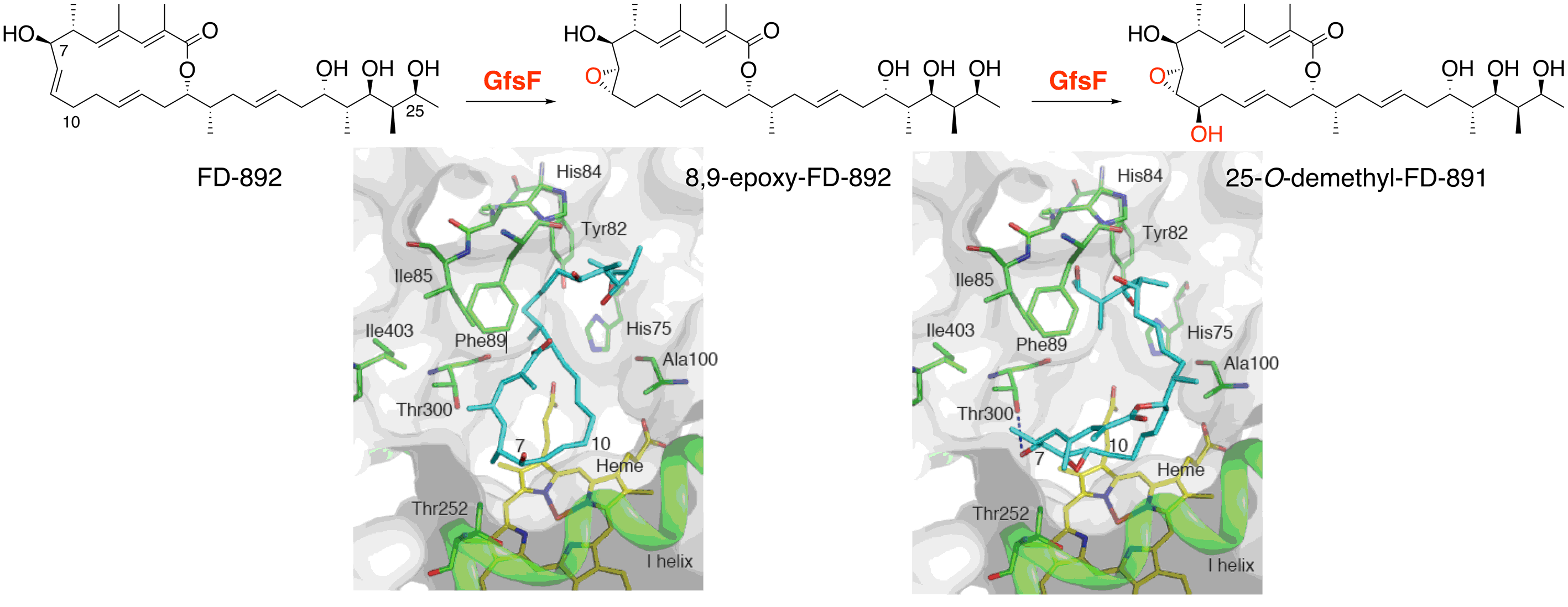
Miyanaga, A.; Takayanagi, R.; Furuya, T.; Kawamata, A.; Itagaki, T.; Iwabuchi, Y.; Kanoh, N.; Kudo, F.; Eguchi, T. Substrate Recognition by a Dual Functional P450 Monooxygenase Involved in FD-891 Biosynthesis. ChemBioChem, 2017, 18, 2179-2187.
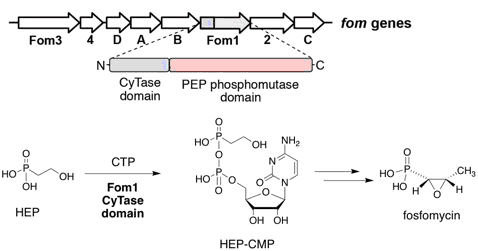
Cho, S.-H.; Kim, S.-Y.; Tomita, T.; Shiraishi,T.; Park, J.-S.; Sato, S.; Kudo, F.; Eguchi,T.; Funa,N.; Nishiyama, M.; Kuzuyama,T. Fosfomycin Biosynthesis via Transient Cytidylylation of 2-Hydroxyethylphosphonate by the Bifunctional Fom1 Enzyme. ACS Chem. Biol., 2017, 12, 2209-2215.
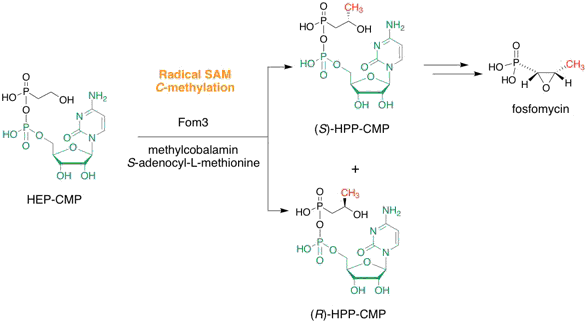
Sato, S.; Kudo, F.; Kim, S.-Y.; Kuzuyama, T.; Eguchi, T. Methylcobalamin-Dependent Radical SAM C-Methyltransferase Fom3 Recognizes Cytidylyl-2-hydroxyethylphosphonate and Catalyzes the Nonstereoselective C-Methylation in Fosfomycin Biosynthesis.
Biochemistry, 2017, 56, 3519-3522.
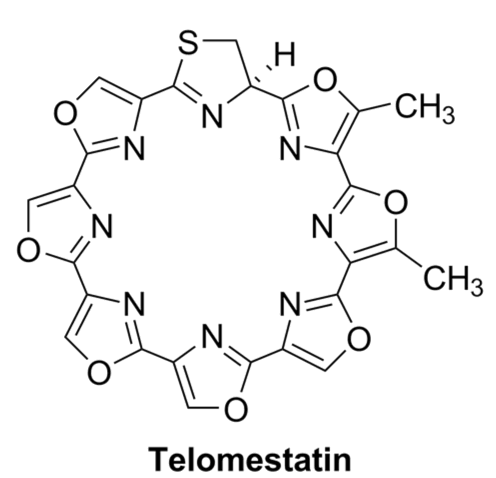
Amagai, K.; Ikeda, H.; Hashimoto, J.; Kozone, I.; Izumikawa, M.; Kudo, F.; Eguchi, T.; Nakamura, T.; Osada, H.; Takahashi, S.; Shin-ya, K. Identification of a Gene Cluster for Telomestatin Biosynthesis and Heterologous Expression Using a Specific Promotor in a Clean Host.
Chisuga, T.; Miyanaga, A.; Kudo, F.; Eguchi, T. Structural Aanalysis of the Dual Function Thioesterase SAV606 Unravels the Mechanism of Michael Addition of Glycine to an α,β-Unsaturated Thioester. J. Biol. Chem., 2017, 292, 10926-10937.
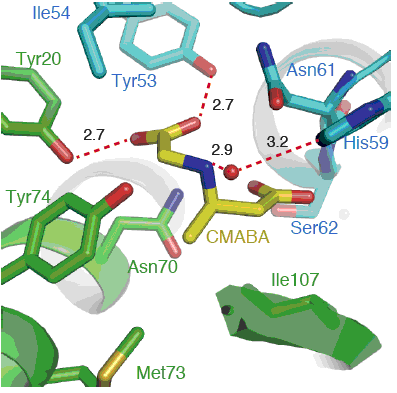
Cieślak, J.; Miyanaga, A.; Takaku, R.; Takaishi, M.; Amagai, K.; Kudo, F.; Eguchi, T. Biochemical Characterization and Structural Insight into Aliphatic β-Amino Acid Adenylation Enzymes IdnL1 and CmiS6. Proteins, 2017, 85, 1238-1247.
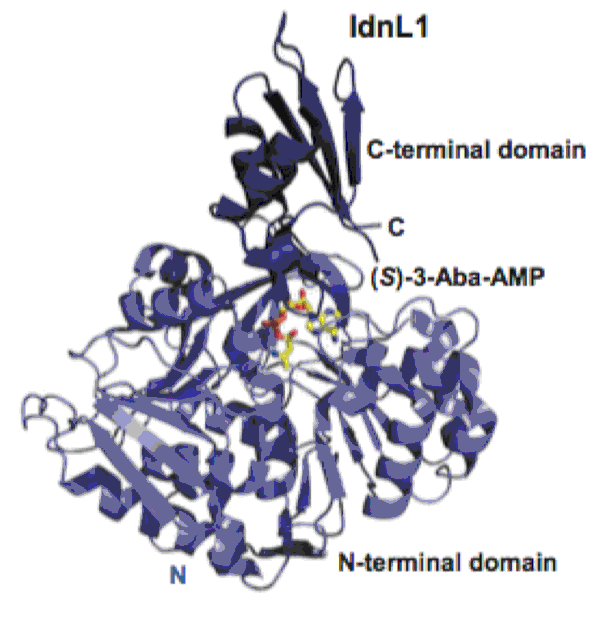
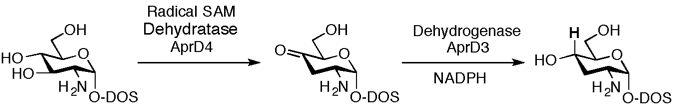
Kudo, F.; Tokumitsu, T.; Eguchi, T. Substrate Specificity of Radical S-Adenosyl-L-methionine Dehydratase AprD4 and Its Partner Reductase AprD3 in the C3′-Deoxygenation of Aminoglycoside Antibiotics. J. Antibiot., 2017, 70, 423-428.
2016
Miyanaga, A.; Kudo, F.; Eguchi, T. Mechanisms of β-Amino Acid incorporation in Polyketide Macrolactam Biosynthesis. Curr. Opin. Chem. Biol., 2016, 35, 58-64.
宮永顕正, 工藤史貴, 江口正, 天然物生合成酵素によるキャリアータンパク質の認識機構, バイオサイエンスとインダストリー, 2016, 74, 382-387.
Kudo, F.; Tsunoda, T.; Takashima, M.; Eguchi, T. Five-membered Cyclitol Phosphate Formation by a myo-Inositol Phosphate Synthase Ortholog in the Biosynthesis of the Carbocyclic Nucleoside Antibiotic Aristeromycin. ChemBioChem, 2016, 17, 2143-2148.
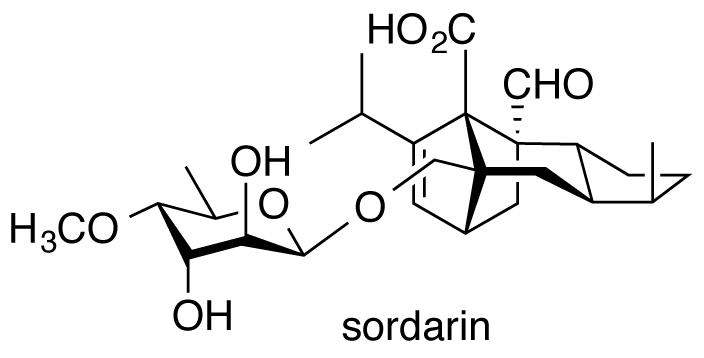
Kudo, F.; Matsuura, Y.; Hayashi, T.; Fukushima, M.; Eguchi, T. Genome Mining of the Sordarin Biosynthetic Gene Cluster from Sordaria araneosa Cain ATCC 36386: Characterization of Cycloaraneosene Synthase and GDP-6-Deoxyaltrose Transferase. J. Antibiot., 2016, 69, 541-548.
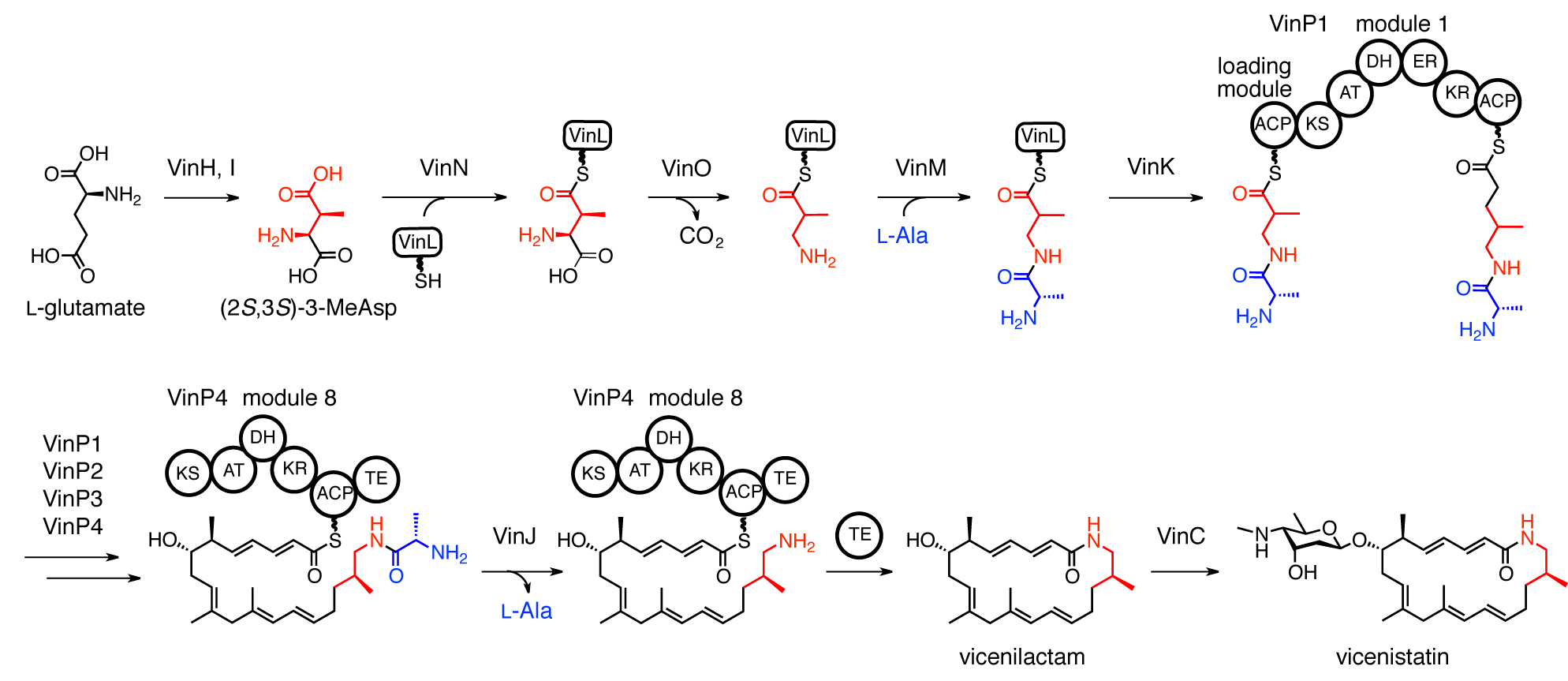
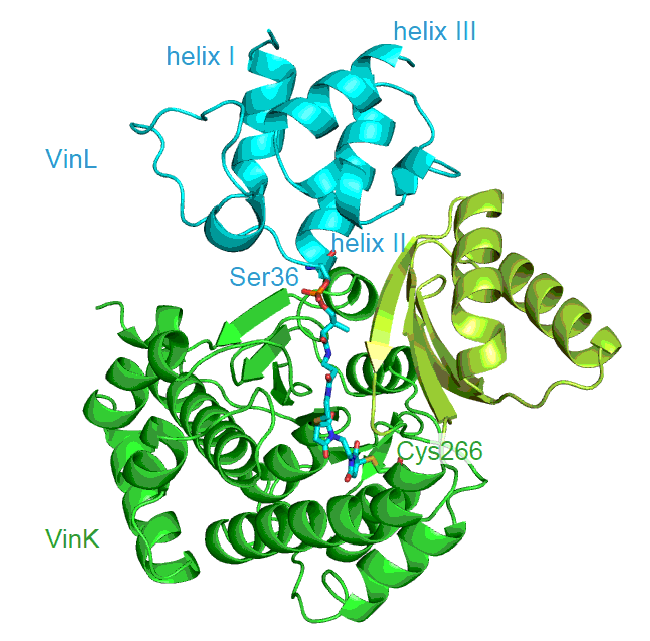
Miyanaga, A.; Iwasawa, S.; Shinohara, Y.; Kudo, F.; Eguchi, T. Structure-based Analysis of the Molecular Interactions between Acyltransferase and Acyl Carrier Protein in Vicenistatin Biosynthesis. Proc. Natl. Acad. Sci. USA, 2016, 113, 1802-1807.
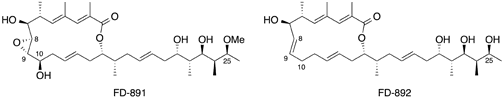
Kanoh, N.; Itagaki, T.; Kawamata, A.; Takeuchi, M.; Hamada, K.; Iwabuchi, Y.; Eguchi,T.; Kudo,F.; Usui,T. Synthesis and Structure-activity Relationship Study of FD891: Importance of the Side Chain and C8-C9 Epoxide for Cytotoxic Activity against Cancer Cells. J. Antibiot., 2016, 69, 287-293.
Nishiyama, Y.; Ohmichi, T.; Kazami, S.; Iwasaki, H.; Mano, K.; Nagumo, Y.; Kudo, F.; Ichikawa, S.; Iwabuchi, Y.; Kanoh, N.; Eguchi, T.; Osada, H.; Usui, T. Vicenistatin Induces the Early Endosome-derived Vacuole Formation in Mammalian Cells. Biosci. Biotechnol. Biochem., 2016, 80, 902-910.
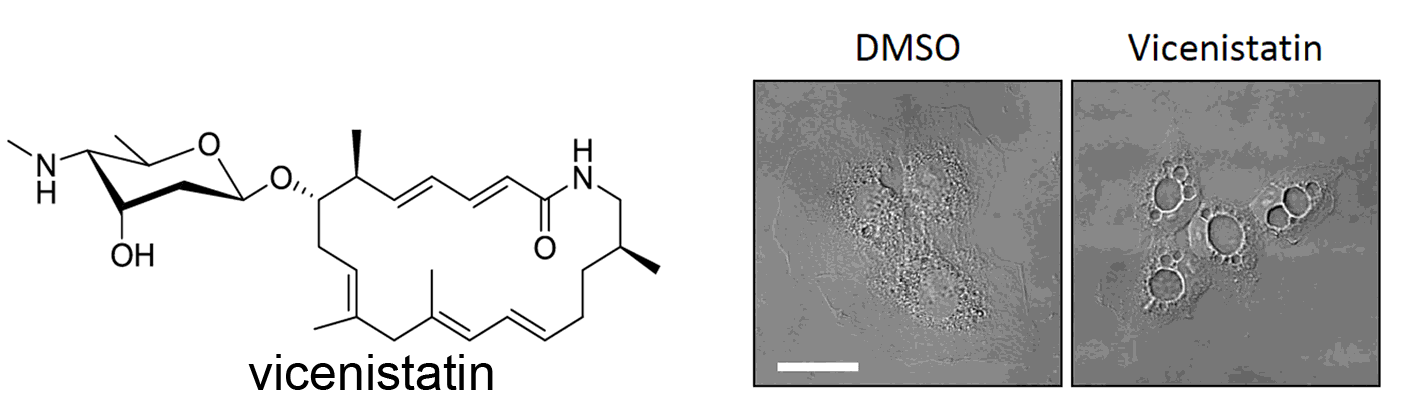
Miyanaga, A.; Hayakawa, Y.; Numakura, M.; Hashimoto, J.; Teruya,K.; Hirano, T.; Shin-ya, K.; Kudo, F.; Eguchi, T. Identification of the Fluvirucin B2 (Sch 38518) Biosynthetic Gene Cluster from Actinomadula fulva subsp. indica ATCC 53714: Substrate Specificity of β-amino Acid Selective Adenylating Enzyme FlvN. Biosci. Biotechnol. Biochem., 2016, 80, 935-941.
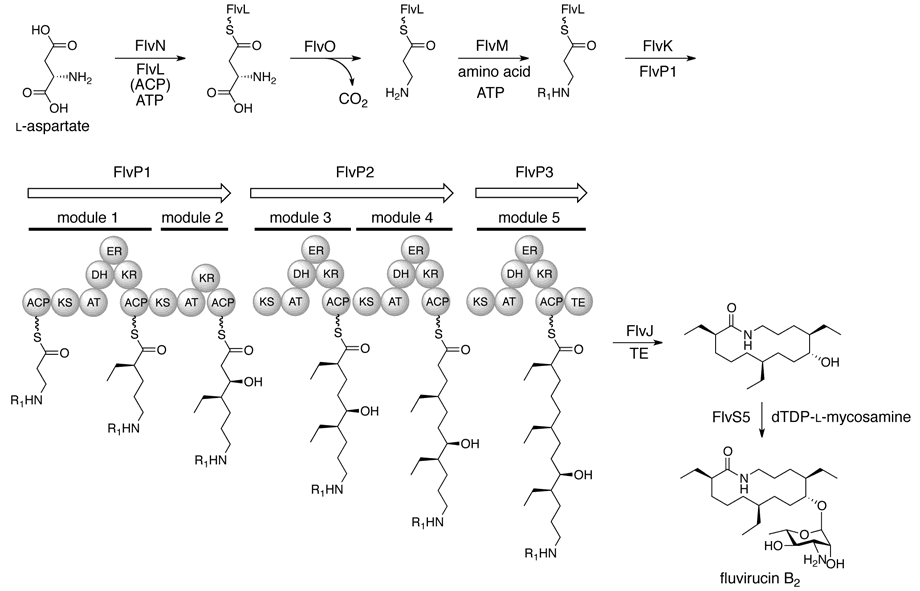
Kudo, F.; Kawamura, K.; Furuya, T.; Yamanishi,H.; Motegi, A.; Komatsubara, A.; Numakura, M.; Miyanaga, A.; Eguchi, T. Parallel Post-Polyketide Synthase Modification Mechanism Involved in FD-891 Biosynthesis in Streptomyces graminofaciens A-8890. ChemBioChem, 2016, 17, 233-238.
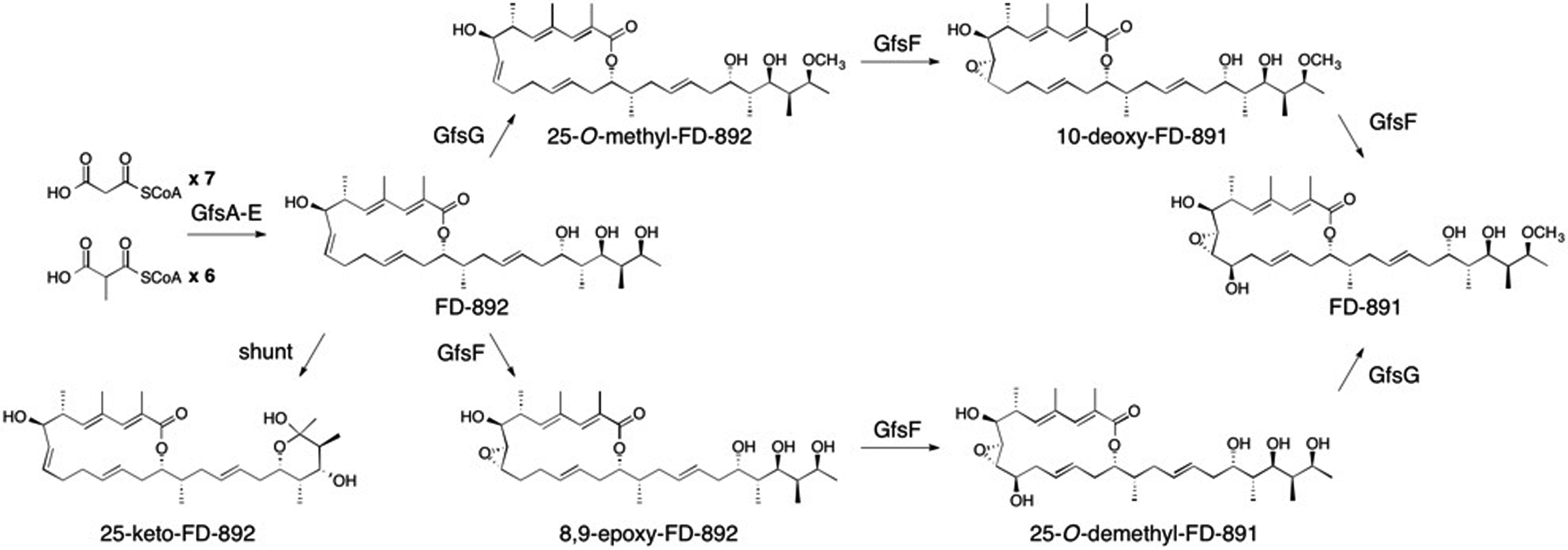
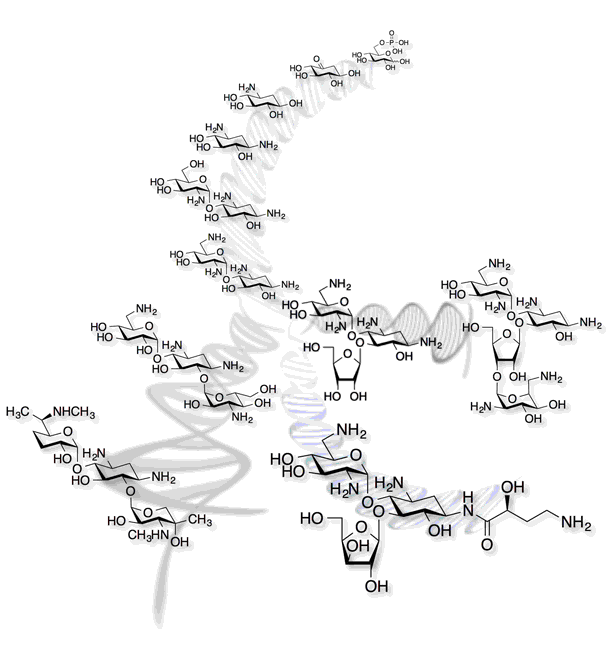
Kudo, F.; Eguchi, T. Aminoglycoside Antibiotics: New Insights into the Biosynthetic Machinary of Old Drugs. Chem. Rec., 2016, 16, 4-18.
2015
Hirayama, A.; Miyanaga, A.; Kudo, F.; Eguchi, T. Mechanism-based Trapping of the Quinonoid Intermediate Using the K276R Mutant of PLP-Dependent 3-Aminobenzoate Synthase PctV in the Biosynthesis of Pactamycin. ChemBioChem, 2015, 16, 2484-2490.
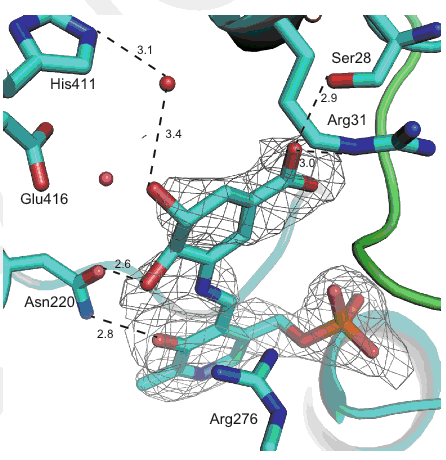
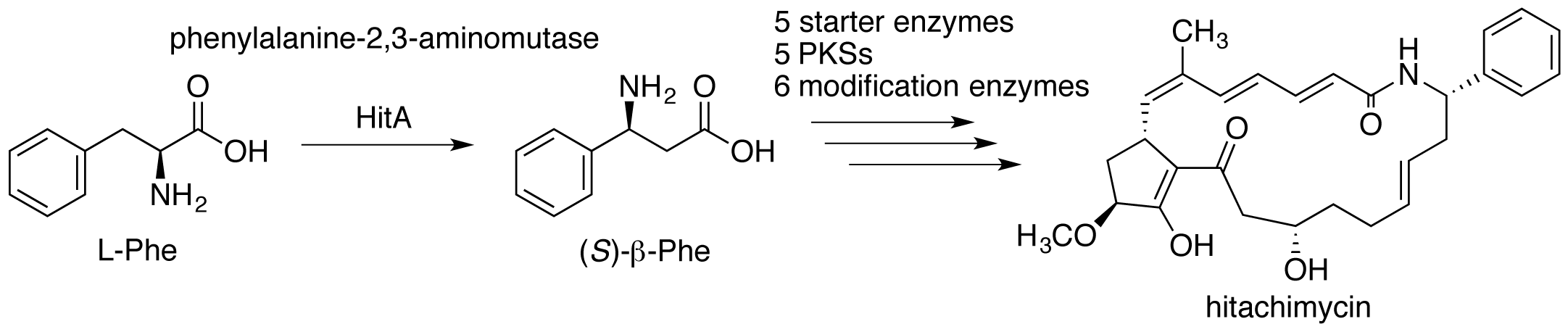
Kudo, F.; Kawamura, K.; Uchino, A.; Miyanaga, A.; Numakura,M.; Takayanagi, R.; Eguchi, T. Genome Mining of Hitachimycin Biosynthetic Gene Cluster; Involvement of Phenylalanine-2,3-aminomutase in the Biosynthesis. ChemBioChem, 2015, 16, 909-914.
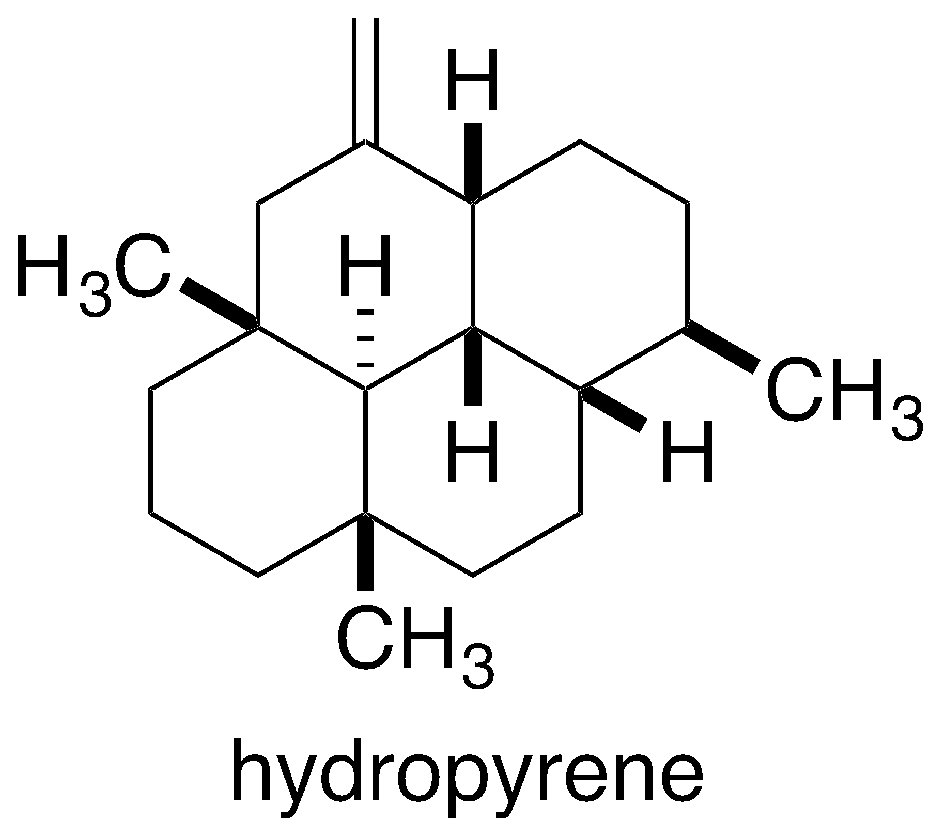
Yamada, Y.; Arima, S.; Nagamitsu, T.; Johmoto, K.; Uekusa, H.; Eguchi, T.; Shin-ya, K.; Cane,D. E.; Ikeda, H. Novel Terpenes Generated by Heterologous Expression of Bacterial Terpene Synthase Genes in an Engineered Streptomyces Host. J. Antibiot., 2015, 68, 385-394.
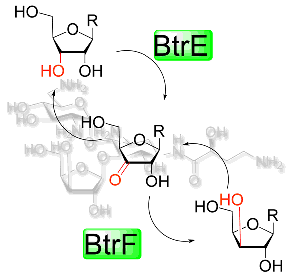
Takeishi, R.; Kudo, F.; Numakura, M.; Eguchi, T. Epimerization at C-3’’ in Butirosin Biosynthesis by an NAD(+)-Dependent Dehydrogenase BtrE and an NADPH-Dependent Reductase BtrF. ChemBioChem, 2015, 16, 487-495.